Publishing extensions
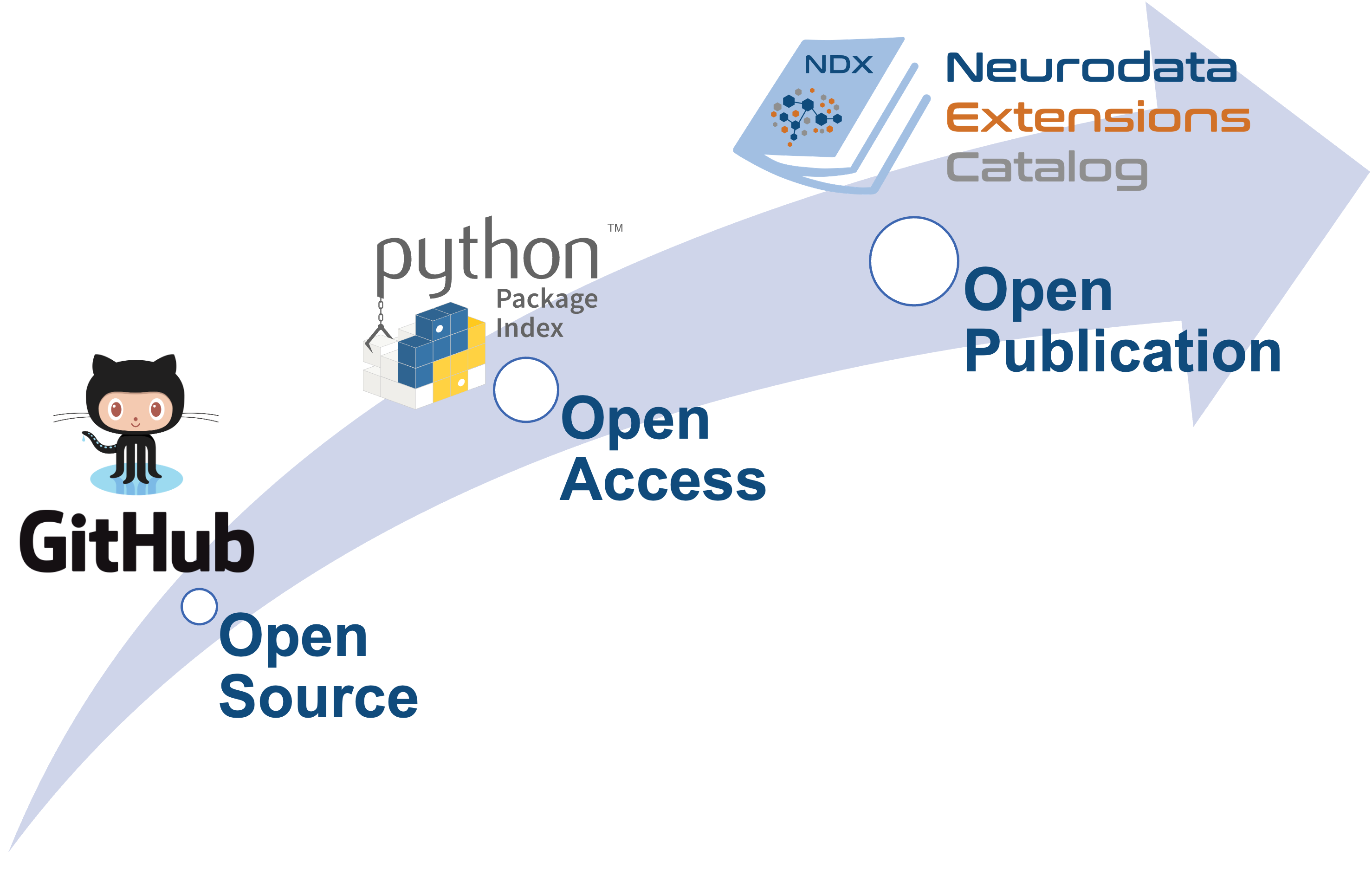
Neurodata extensions can be shared with the community using the NDX Catalog. As illustrated in the figure, the publication process is divided into three main steps: 1) open release of the sources to the community using GitHub, 2) open access of versioned releases via PyPI, and 3) open publication of the extension to the community via the NDX Catalog.
Open Source: Releasing your extension Git repository
Before publishing your extension online you should add a license file. Permissive licenses should be used if possible. A BSD license <https://opensource.org/licenses/BSD-3-Clause> is recommended.
Modify
README.mdat the root directory of your extension repo to describe the extension for interested developers and users.The first step to publishing your extension then is to make your Git repository accessible online via GitHub, or any other public Git hosting service of your choice. To publish your extension on GitHub you will need a GitHub account and follow the following instructions to add an existing project to GitHub
Make a release for the extension on GitHub with the version number specified. e.g. if the version is
0.1.0, then this page should exist:https://github.com/<my_username>/<my_extension>/releases/tag/0.1.0. See the creating a release guide on GitHub for instructions on how to make a release. See the NWB Versioning Guidelines for details on how to version extensions.
Note
We here focus on GitHub because it is the services that is currently most commonly used for extensions repositories. However, users may chose to use other services (e.g., GitLab or Bitbucket) to share their sources.
Open Access: Releasing your extension on PyPI
To make your extension installable for users via pip and manage public releases NWB uses the Python Package Index (PyPI) index.
Follow these directions to package your project. You may need to modify
setup.py. If your extension version is0.1.0, then this page should exist:https://pypi.org/project/<myextension>/0.1.0Once your GitHub release and
setup.pyare ready, publishing on PyPI:
python setup.py sdist bdist_wheel
twine upload dist/*
Open Publication: Publishing your extension on the NDX Catalog
The NDX Catalog serves as a central, community-led catalog for extensions to the NWB data standard. The NDX Catalog manages basic metadata about extensions while ownership of the source repositories for the extensions remain with the developers. To publish your extension on the catalog:
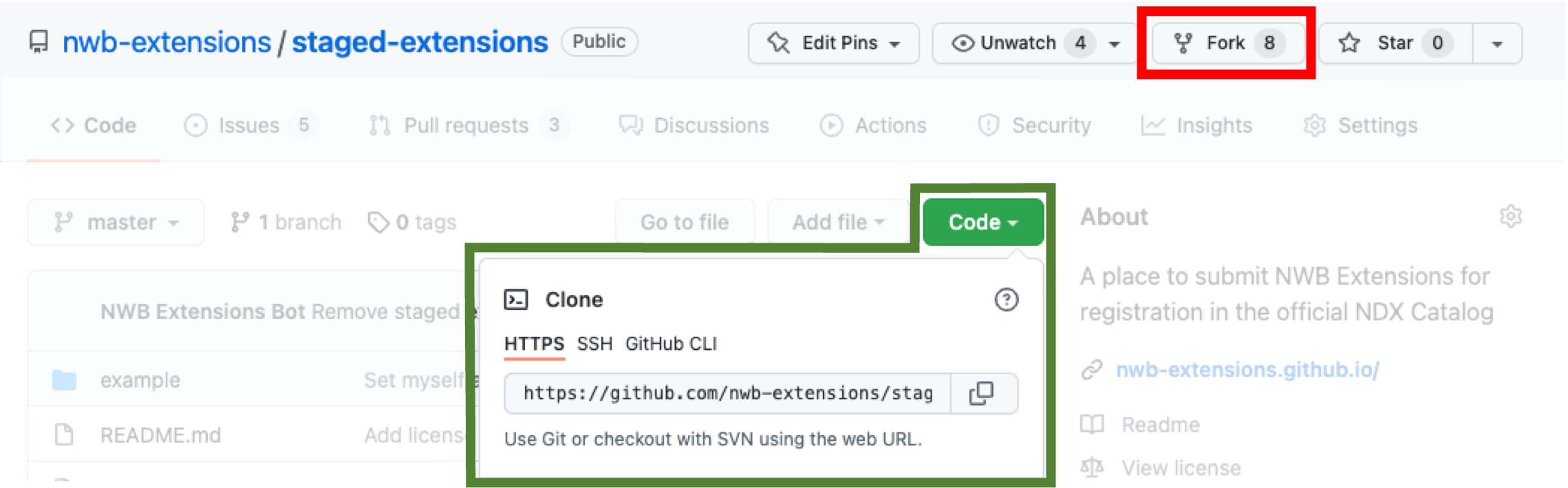
Fork the staged-extensions repository, which is used to submit new extension to the catalog via pull requests.
Clone your fork of the staged-extensions onto your computer, e.g., via
git clone <my_fork_url>
Copy the directory
staged-extensions/exampleto a new directory with the name of your extension, e.g., viacp -r staged-extensions/example staged-extensions/<my_extension>Edit
staged-extensions/<my_extension>/ndx-meta.yamlwith information on where to find your NWB extension. TheNEXTSTEPS.mdfile in thendx-templateincludes an autogenerated templatendx-meta.yamlfile that you may copy and modify. The YAML file MUST contain a dict with the following keys:name: extension namespace nameversion: extension versionsrc: URL for the main page of the public repository (e.g. on GitHub, BitBucket, GitLab) that contains the sources of the extensionpip: URL for the main page of the extension on PyPIlicense: name of the license of the extension- `` maintainers``: list of GitHub usernames of those who will reliably maintain the extension
You may copy and modify the following YAML that was auto-generated:
Edit
staged-extensions/<my_extension>/README.mdto add information about your extension. Usually, you can here just copy theREADME.mdfrom your extension repocp <my_extension>/README.md staged-extensions/<my_extension>/README.mdAdd and commit your changes to Git and push your changes to GitHub:
cd staged-extensions
git add <my_extension>
git commit -m "Add new catalog entry for <my_extension>"
git push
Open a pull request. See the creating a pull request from a fork website for step-by-step instructions on to create a pull request on GitHub.
Once the PR has been created, building of your extension will be tested on Windows, Mac, and Linux. The technical team will review your extension shortly after and provide feedback and request changes, if any. Once the technical team has approved and merged your pull request, a new repository, called
<my_extension>-recordwill be created in the nwb-extensions GitHub organization and you will be added as a maintainer for that repository.
Updating your published extension
Once you have published your extension you can update and publish new version as follows:
Update your
<my_extension>GitHub repositoryPublish your updated extension on PyPI.
Fork the
<my_extension>-recordrepository from the nwb-extensions GitHub organization and update yourndx-meta.yaml,README.mdand other relevant record dataOpen a pull request to test the changes automatically.
The technical team will review your changes shortly after and provide feedback and request changes, if any.
Your updated extension is ready once your PR has been approved and merged.
Policies: Neurodata Extension (NDX) rules and guidelines
Sharing Guidelines: requirements and strategy for sharing format extensions for NWB
Sharing Strategies: standard practices and strategies for sharing format extensions for NWB
Proposal Review Process: process by which extensions to the NWB core standard are proposed, evaluated, reviewed, and accepted
Versioning Guidelines: requirements and strategy for versioning namespaces for the NWB core schema and extensions